This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Introduction
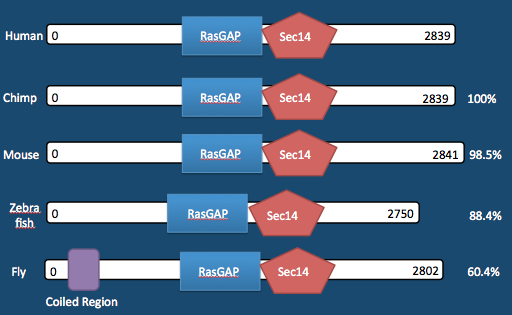
Neurofibromatosis Type 1 (NF1) is one of the most prevalent inherited genetic neurological disorders in the world affecting one in 3,000 births. Neurofibromatosis is characterized by tumor growth along the nervous system, including the brain, spinal cord, nerves and skin. Around half of patients with NF1 also have learning disabilities with largely varying phenotypes. NF1 is caused by an autosomal dominant mutation anywhere along a single gene, the Nf1 gene, which encodes for the protein neurofibromin (NF1). Neurofibromin is a 2839 amino acid long protein with a RasGAP domain in the middle. NF1 is typically found in the nucleus, membrane, or cytoplasm of axons and dendrites, it acts as a GTPase activator along with other binding abilities. The biological processes that NF1 acts in are numerous including: cell communication, negative regulation of Ras, heart/liver/forebrain/cerebral cortex development, observational learning, MAPK cascade, myelination, etc.
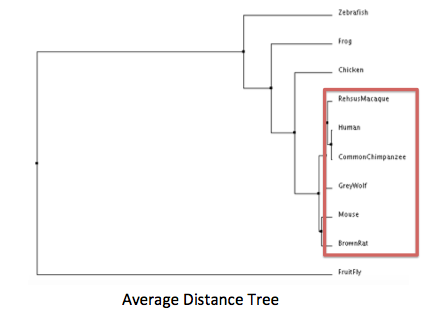
NF1 is a highly conserved protein across mammals as shown in Figure 1,2. Zebrafish’s neurofibromin percent identity to humans drops down to 88.4, and fruitfly to 60.4. (See pages Homology, Phylogeny, Domains).
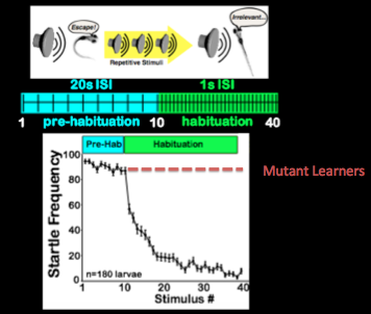
My main goal is to understand how NF1 regulates learning through cellular and molecular components. I am going to use zebrafish to do this because they have a defined assay for measuring learning ability, have a defined and functional neural circuit at 5 days post fertilization that can be easily imaged, and have a short life cycle.
My main goal is to understand how NF1 regulates learning through cellular and molecular components. I am going to use zebrafish to do this because they have a defined assay for measuring learning ability, have a defined and functional neural circuit at 5 days post fertilization that can be easily imaged, and have a short life cycle.
Specific Aims
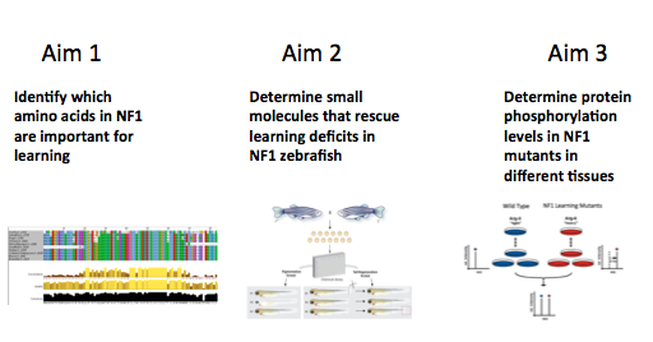
Aim 1: Identify conserved amino acids in NF1 protein necessary for learning.
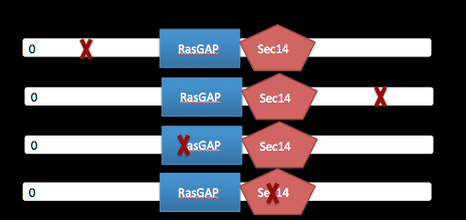
Amino acids that are conserved in both simple learning and advanced learning organisms, and those that are specific to advanced learning organisms, will be important to understand how NF1 functions in the learning process. The NF1 protein is highly conserved across species with advanced learning (humans, chimps, gorillas, etc.) and simple learning (i.e. mice, rats, zebrafish, frog, and fruit fly). To determine which amino acids are important for learning, Clustal Omega will be used to align amino acid sequences (shown in Phylogeny page) and analyze the areas that are differing in polarity or subgroup of amino acid between advanced learners and simple learners. Once candidate amino acids have been identified, transgenic knockout zebrafish will be created using CRISPER. Then the fish will be assayed for learning ability to identify the conserved amino acids important for learning. Identifying key amino acids important for learning will lead to a better understanding of how mutations in NF1 lead to learning disabilities.
Aim 2: Identify small molecules that rescue learning deficits in NF1 zebrafish.
Learning function will be rescued in Nf1 mutants by certain small molecule(s) associated with the cAMP pathway. I chose to look at the cAMP pathway instead of the Ras pathway because if given Ras inhibitors, NF1 patients did not recover from learning disabilities, so another pathway that NF1 effects is what I am going to look at. Transgenic zebrafish NF1 learning mutants from Aim1 will be used in a phenotype-based forward chemical genetic screen. A focused chemical library will be developed to screen the small molecules associated with the cAMP pathway. Our phenotype of interest would be rescued function of learning. Controls will be the percentage of wild type fish that preform the learning assays. Small molecules that rescue Nf1 mutants with learning disabilities will lead to a better understanding of signaling pathways that NF1 acts through in the brain.
Aim 3: Determine protein phosphorylation levels in NF1 mutants in different tissues.
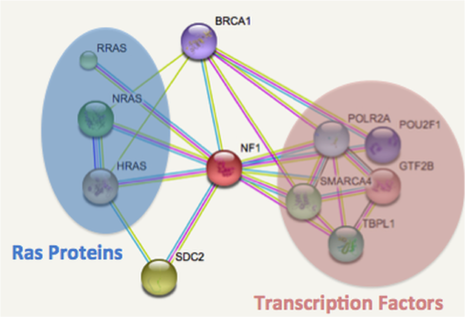
In Nf1 learning mutants there will be lower phosphorylation levels in the cAMP pathway’s proteins due to inactivation by the mutant NF1 protein. Quantitative phosphorylation with mass spectrometry will be completed on the learning mutant Nf1 zebrafish in neuronal tissue vs. other tissue types. Proteins with differing phosphorylation levels in the neuronal tissue vs. other tissues will be analyzed for what pathway they participate in through STRING interaction networks to determine if proteins associated with part of the cAMP pathway are affected in neuronal tissue. Phosphorylation levels during disease state like NF1 can illuminate which pathways are not functioning.
Aim 3: Determine protein phosphorylation levels in NF1 mutants in different tissues.
In Nf1 learning mutants there will be lower phosphorylation levels in the cAMP pathway’s proteins due to inactivation by the mutant NF1 protein. Quantitative phosphorylation with mass spectrometry will be completed on the learning mutant Nf1 zebrafish in neuronal tissue vs. other tissue types. Proteins with differing phosphorylation levels in the neuronal tissue vs. other tissues will be analyzed for what pathway they participate in through STRING interaction networks to determine if proteins associated with part of the cAMP pathway are affected in neuronal tissue. Phosphorylation levels during disease state like NF1 can illuminate which pathways are not functioning.
Future Directions
With the predicted results from the above experiments we can move forward in the future to make predictions on the severity of learning disabilities correlating to the position of the mutation on the NF1 gene. With this ability diagnosis and treatments can be tailored to an individual with NF1 to better deal with learning disabilities presenting in NF1 patients.
With the identified small molecules from the chemical drug screen in Aim2 we could proceed with drug trials for mice then humans for those small molecules that are rescuing learning disabilities in NF1 zebrafish.
With the identified small molecules from the chemical drug screen in Aim2 we could proceed with drug trials for mice then humans for those small molecules that are rescuing learning disabilities in NF1 zebrafish.
Presentation Rough Draft 1
| johnson_presentation_draft_1.pptx |
Presentation Rough Draft 2
| final_talks_rough_draft_2_040417_2.zip |
Presentation Final Draft
| final_talks_final_draft.pdf |