Proteomics
This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
A proteome is the collection of all the proteins in an organism. This can be narrowed down to different sections of the organism such as cell type or tissue type. The proteome can also be analyzed at different time points. The proteome of a given organism will be constantly changing in response to its environment and biological needs. Proteomics looks at which proteins are interacting with each other under certain biological conditions. [1]
Proteins are often post-translationally modified to be able to function in their proper way. Common post translational modifications are: phosphorylation, ubiquitination, methylation, acetylation, etc. [1]
A common way for scientists to be alerted to the fact that a signaling pathway is functioning at a given time in a cell is to look at the protein phosphorylation in the cell type. If proteins are phosphorylated their signaling pathway has most likely been activated. [1]
Proteins are often post-translationally modified to be able to function in their proper way. Common post translational modifications are: phosphorylation, ubiquitination, methylation, acetylation, etc. [1]
A common way for scientists to be alerted to the fact that a signaling pathway is functioning at a given time in a cell is to look at the protein phosphorylation in the cell type. If proteins are phosphorylated their signaling pathway has most likely been activated. [1]
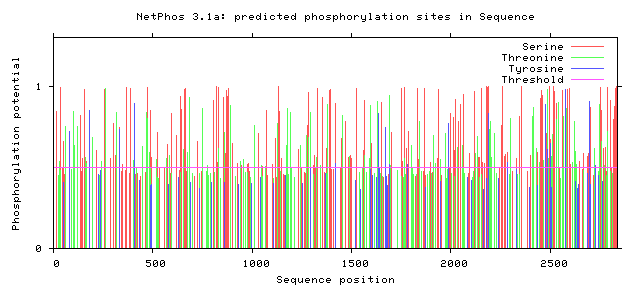
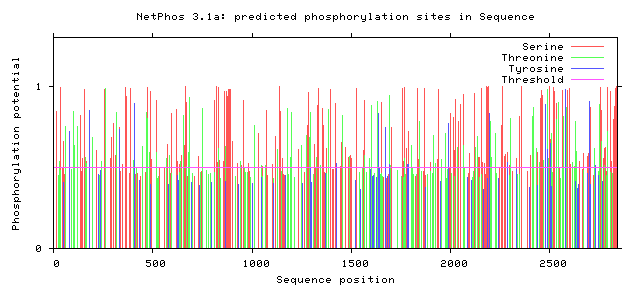
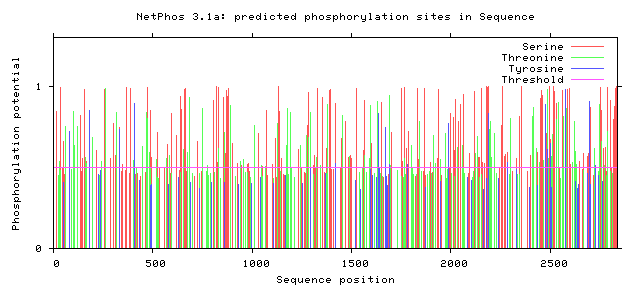
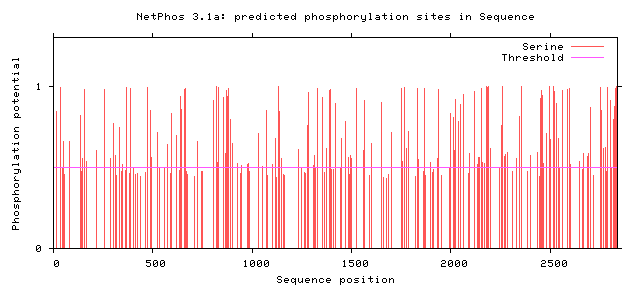
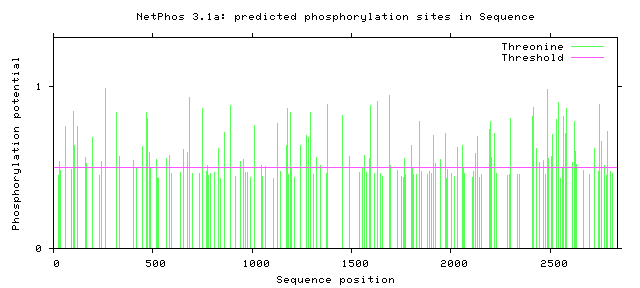
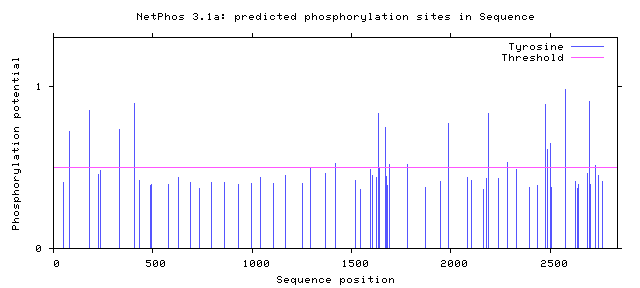
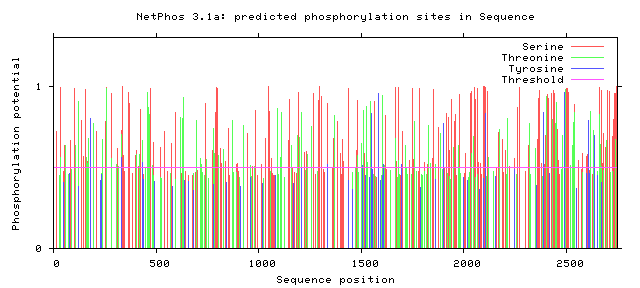
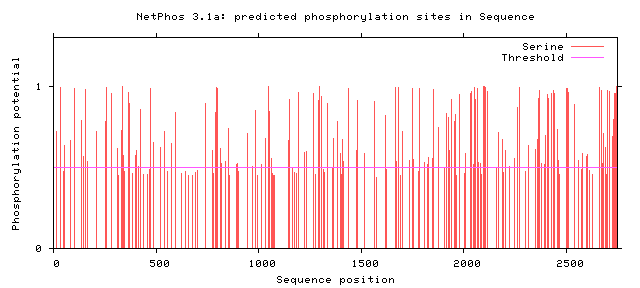
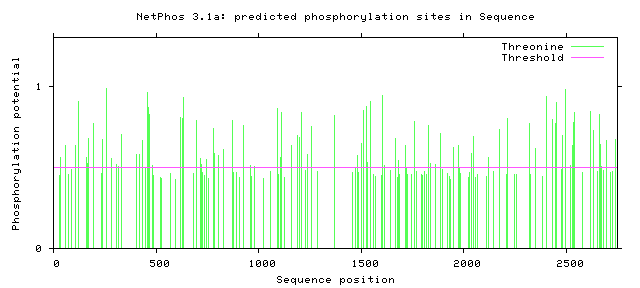
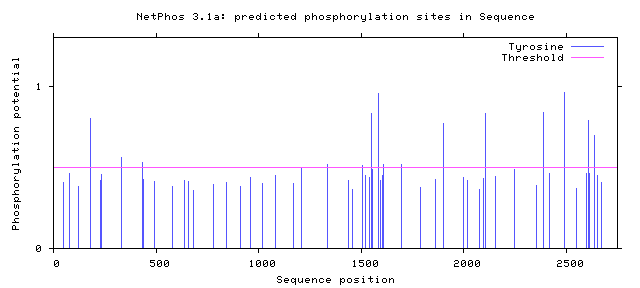
NF1 Phosphorylation sites
Human NF1 phosphorylation sites
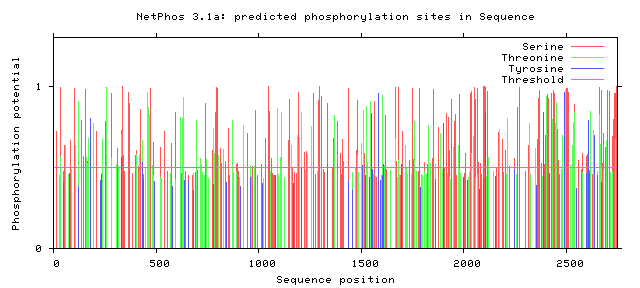
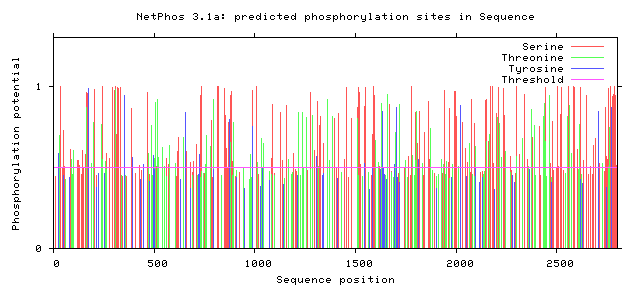
Zebrafish NF1 phosphorylation sites
References
All Phosphorylation Figures: Net Phos.
[1] Proteomics. Wikipedia. https://en.wikipedia.org/wiki/Proteomics
[1] Proteomics. Wikipedia. https://en.wikipedia.org/wiki/Proteomics